BioGRID([path, file_resources, …])
|
|
BioMart([host, verbose, cache, secure])
|
Interface to the BioMart service |
BioMartManager(dataset, attributes, host, …)
|
|
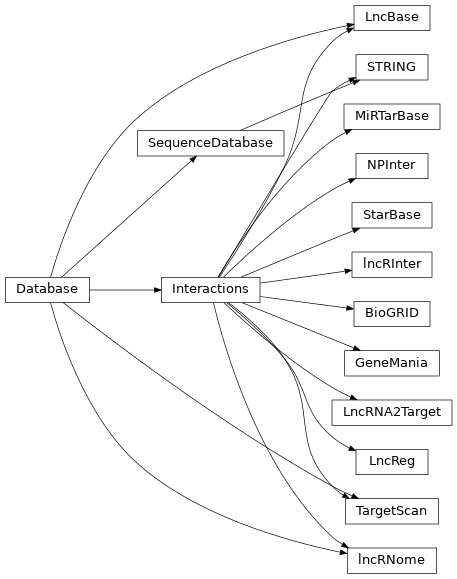
Database(path[, file_resources, col_rename, …])
|
This is a base class used to instantiate an external Database given a a set of files from either local files or URLs. |
Dict(*args, **kwds)
|
|
EnsemblGeneSequences([biomart, attributes, …])
|
|
EnsemblGenes([biomart, attributes, host, …])
|
|
EnsemblSNP([biomart, attributes, host, …])
|
|
EnsemblSomaticVariation([biomart, …])
|
|
EnsemblTranscriptSequences([biomart, …])
|
|
GTEx([path, file_resources, col_rename, …])
|
|
GeneMania(path[, file_resources, …])
|
|
Interactions(path, file_resources[, …])
|
|
List(*args, **kwds)
|
|
LncBase(path[, file_resources, …])
|
|
LncRNA2Target([path, file_resources, …])
|
|
LncReg(path, file_resources[, …])
|
|
MiRTarBase([path, file_resources, …])
|
|
NONCODE(path[, file_resources, col_rename, …])
|
|
NPInter([path, file_resources, …])
|
|
ProteinAtlas([path, file_resources, …])
|
|
RNAcentral([path, file_resources, …])
|
|
STRING([path, file_resources, species_id, …])
|
|
SequenceDatabase([replace_U2T])
|
Provides a series of methods to extract sequence data from SequenceDataset. |
StarBase(path, file_resources[, …])
|
|
StringIO([initial_value, newline])
|
Text I/O implementation using an in-memory buffer. |
TANRIC(path[, file_resources, col_rename, …])
|
|
TargetScan(path[, file_resources, …])
|
|
lncRInter(path[, file_resources, …])
|
|
lncRNome(path, file_resources[, …])
|
|